Home - nsalomonis/altanalyze GitHub Wiki
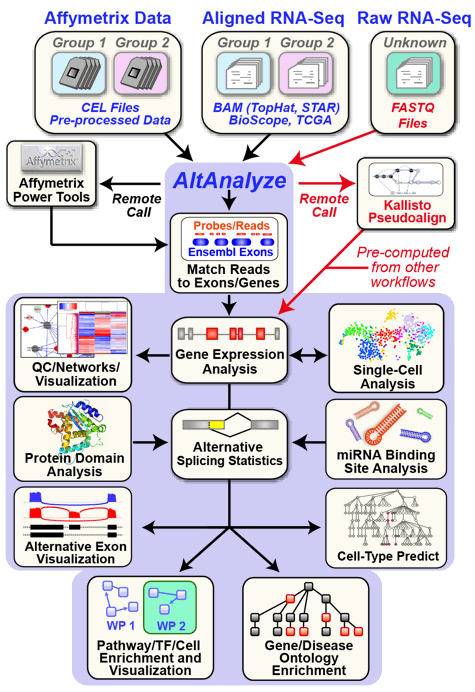
AltAnalyze is an extremely user-friendly and open-source toolkit that can be used for a broad range of genomics analyses. These analyses include the direct processing of raw RNASeq or microarray data files, advanced methods for single-cell population discovery with ICGS, differential expression analyses, analysis of alternative splicing/promoter/polyadenylation and advanced isoform function prediction analysis (protein, domain and microRNA targeting). Multiple advanced visualization tools and à la carte analysis methods are supported in AltAnalyze (e.g., network, pathway, splicing graph). AltAnalyze is compatible with various data inputs for RNASeq data (FASTQ, BAM, BED), microarray platforms (Gene 1.0, Exon 1.0, junction and 3' arrays) to enable direct analysis of gene expression and alternative splicing. This software requires no advanced knowledge of bioinformatics programs or scripting or advanced computer hardware. User friendly videos, online tutorials and blog posts are also available. Command-line examples can be found here.
AltAnalyze documentation can be found on Read The Docs and stand-alone archives are provided at sourceforge as well as at GitHub. For questions not addressed here, please contact us.
News Update 12/16/18