MAGeT Processing - katielavigne/documentation GitHub Wiki
Preamble
We primarily use MAGeTbrain to measure brain volumes, particularly of the hippocampus, its subfields, and adjacent areas. The first step is to select 21 subjects from your sample to use as templates. These subjects should all have good quality T1-weighted (and T2-weighted, if available) scans and they should be representative of your sample (approximately half controls/patients, half males/females, and similar age distribution to your sample). MAGeT is run on Niagara using BPIPE processed data. If you have T2-weighted images, it is preferable to use those as they provide better visualization of subcortical areas.
Setting up MAGeT
Once you've logged into Niagara, create a MAGeT directory in your scratch:
mkdir -p ${SCRATCH}/maget
Load required modules:
module load cobralab
Clone the main MAGeT branch:
git clone -b simplified-labelmask https://github.com/CobraLab/MAGeTbrain.git $SCRATCH/MAGeTbrain
Note: If you are interested in additional morphometric measures (displacement, surface area), you can instead clone the MAGeT morpho branch):
git clone -b morpho-labelmask https://github.com/CobraLab/MAGeTbrain.git. Switch between branches by tracking the library you didn't clone:git checkout -track simplified-labelmask. Once both are tracked, switch:git checkout simplified-labelmask.
Source MAGeTbrain:
source $SCRATCH/MAGeTbrain/bin/activate
Change directory to the maget directory you created:
cd $SCRATCH/maget
Initialize the MAGeTbrain directory structure:
mb init
This will create an input folder with a full MAGeTbrain directory tree.
Setting up your data
You must then copy (cp) all your files to the relevant directories just created.
Copy BEaST extracted files to input/subjects/brains:
- T1-weighted:
find ${SCRATCH}/bpipe/t1w -name *"cutneckapplyautocrop.beastextract.mnc" -exec cp {} ${SCRATCH}/maget/input/subjects/brains \;
- T2-weighted:
find $SCRATCH/t2-corrected/ -name *"T2_corrected.mnc" -exec cp {} ${SCRATCH}/maget/input/subjects/brains \;
Copy template files to templates/brains directory. First, create a template_subjects.txt file and enter your 21 template subject IDs on separate lines. These subject IDs must correspond to the original BPIPE input filenames, excluding the file extension (.mnc):
nano $SCRATCH/maget/template_subjects.txt
 Save (Ctrl+o, then Enter) and close (Ctrl+x) the file.
Save (Ctrl+o, then Enter) and close (Ctrl+x) the file.
Read text file and copy images to relevant directories:
- T1-weighted:
while read line; do cp $SCRATCH/maget/input/subjects/brains/${line}.convert.n4correct.cutneckapplyautocrop.beastextract.mnc ${SCRATCH}/maget/input/templates/brains/; done < $SCRATCH/maget/template_subjects.txt
- T2-weighted:
while read line; do cp $SCRATCH/t2-corrected/${line}_T2_corrected.mnc ${SCRATCH}/maget/input/templates/brains/; done < ${SCRATCH}/maget/template_subjects.txt
Copy atlas labels to atlases directory (this example is for the hippocampus-white-matter atlas):
cp /project/m/mchakrav/atlases_from_git/hippocampus-whitematter/labels/brain?_labels.mnc ${SCRATCH}/maget/input/atlases/labels
Copy atlas images to atlases directory:
- T1-weighted:
cp /project/m/mchakrav/atlases_from_git/T1_extracted/brain?.mnc ${SCRATCH}/maget/input/atlases/brains/
- T2-weighted:
cp /project/m/mchakrav/atlases_from_git/T2_extracted/brain?_t2.mnc ${SCRATCH}/maget/input/atlases/brains/
rename _t2.mnc .mnc input/atlases/brains/*
Note. If using morpho branch, copy model files to model directories:
cp /project/m/mchakrav/atlases-morpho/morpho_models/hc_amyg_cerebellum/model/brains/model_extracted.mnc ${SCRATCH}/maget/input/model/brains/
cp /project/m/mchakrav/atlases-morpho/morpho_models/hc_amyg_cerebellum/model/objects_hc/* ${SCRATCH}/maget/input/model/objects/
cp /project/m/mchakrav/atlases-morpho/morpho_models/hc_amyg_cerebellum/model/xfms/* ${SCRATCH}/maget/input/model/xfms/
Run MAGeT
Check if directories are well-populated (no output = good):
mb check
Check status (this will take some time):
mb status
You should see something like this:
5 atlases, 21 templates, 295 subjects found
105 atlas-to-template registration commands left
6195 template-to-subject registration commands left
30975 transform merging commands left
30975 label propagation commands left
295 label fusion commands left
Or if using morpho branch:
1 models, 5 atlases, 21 templates, 328 subjects found
350 model prep commands left
349 preprocessing of images commands left
105 atlas-to-template registration commands left
2266152 template-to-subject registration commands left
34440 transform merging commands left
34440 label propagation commands left
328 label fusion commands left
103320 model to subj transform merging commands left
68880 subject grid commands left
328 grid average calculation commands left
656 model object displacement calculation commands left
656 displacement blurring commands left
Run MAGeT:
mb run
Note: If you are running morpho, use this to run MAGeT morpho:
mb run --surfaces –voronoi
Note: If you are working with high-resolution data (< 1mm3), you will need to modify the command, e.g.:
mb run --stage-voting-procs 1 --stage-voting-walltime 24:00:00. For LAM high resolution T2-weighted images, the following command works:
mb run --stage-templatelib-walltime 24:00:00 --stage-templatelib-procs 2 --stage-voting-procs 1 --stage-voting-walltime 24:00:00
When MAGeT is complete, you should run mb status again to make sure everything completed.
If you get failed notifications due to for example, timeout or memory issues, you can modify the settings by increasing the walltime (time allowed to run) or the number of processes running (parallelization) for a given stage. MAGeT defaults can be found in the mb script (MAGeTbrain/bin/mb). Current defaults are:
STAGE_REG_ATLAS (atlas-to-template registration): {'procs': 20, 'walltime': '6:00:00'},
STAGE_REG_TEMPL (template-to-subject registration): {'procs': None, 'walltime': '4:00:00'},
STAGE_XFM_JOIN (transform merging): {'procs': None, 'walltime': '0:10:00'},
STAGE_RESAMPLE (label propagation): {'procs': None, 'walltime': '1:00:00'},
STAGE_VOTE (label fusion): {'procs': 5, 'walltime': '12:00:00'},
STAGE_TAR (tar intermediate files): {'procs': 8, 'walltime': '1:00:00'}}
Issues usually occur at the REG_ATLAS and VOTE stages. Examples of how to modify:
mb run --stage-templatelib-walltime 24:00:00 --stage-templatelib-procs 2
mb --save run -j 3 –stage-voting-procs 3 --stage-voting-walltime 24:00:00
You can check on your job with squeue -u $USER or sq. Once preprocessing is complete (may take a few days), your files will be in the output directory.
Acquiring volumes
Volumes can be converted to .csv as follows:
collect_volumes.sh ${SCRATCH}/maget/output/fusion/majority_vote/* > volumes.csv
Then use scp or rsync to copy it to your machine.
Note: If you've run MAGeT morpho, the displacement & surface area files are the "*5mm_blur.txt" files (see instructions). The numbered headers in the output files correspond to those described in the relevant atlases repository.
MAGeT backup
Assuming you are using your $SCRATCH directory to run MAGeT processing, you will need to backup your results somewhere else, since $SCRATCH is purged every 60 days. You can use rsync or scp to copy the results to another remote location (e.g., CIC or your computer).
The most important files to backup are those you want to use for analysis (e.g., volumes.csv or the displacement/new_lefthc/etc .txt files), but you should also backup the labels that MAGeT outputs, which are used to get these values, for further processing (e.g., quantitative T1) and for QC. You should also at least backup a list of your input and template files so you have a record of which ones were used in the analysis.
Here are some example commands to backup the relevant MAGeT files. First, in your Niagara MAGeT directory, create lists for input and template brains:
ls ${SCRATCH}/maget/input/subjects/brains/ > ${SCRATCH}/maget/input_subject_brains.txt
ls ${SCRATCH}/maget/input/templates/brains > ${SCRATCH}/maget/input_templates_brains.txt
Then, login to the remote server where you would like to backup your files and rsync the relevant output directories:
rsync -rv –-include /input/*.txt –-include /output/fusion/ --include /logs/ [email protected]:/path/to/niagara/maget_dir/* path/to/remote/dir/
If you want to backup the atlas-to-template registrations to use again, be sure to backup the MAGeTbrain repository used, as well as the output/registrations and output/templatemasks folders.
rsync -rv –-include /input/*.txt –-include /output/fusion/ --include /logs/ --include /MAGeTbrain --include /output/registrations --include /output/templatemasks [email protected]:/path/to/niagara/maget_dir/* path/to/remote/dir/
You should also note the MAGeTbrain version used:
cd /path/to/MAGeTbrain
git log
Copy the top few lines into a README.txt file and add any additional information relevant to your study, e.g:
MAGeT run for subjects up to ABC_167 and ABC_250.
MAGeTBrain version info:
commit cc3d16d9df8511bb9c3d2425b38a95183061d2c4 (HEAD -> simplified-labelmask, origin/simplified-labelmask, origin/HEAD)
Author: Gabriel A. Devenyi <email redacted>
Date: Tue Mar 23 10:30:26 2021 -0400
Backed up output/templatemasks and output/registrations so that future runs of MAGeT with new subjects can copy these directories into a new MAGeT directory and skip atlas-to-template registration step.
Templates are as follows: ABC_101_T1, ABC_105_T1, ABC_109_T1, ABC_121_T1, ABC_127_T1, ABC_133_T1, ABC_139_T1, ABC_145_T1, ABC_152_T1, ABC_155_T1, ABC_167_T1, ABC_202_T1, ABC_206_T1, ABC_209_T1, ABC_211_T1, ABC_223_T1, ABC_227_T1, ABC_229_T1, ABC_235_T1, ABC_238_T1, ABC_250_T1
Atlases were copied into ${SCRATCH}/maget/input/atlases/{brains,labels} with following commands:
# copy labels
cp /project/m/mchakrav/atlases_from_git/hippocampus-whitematter/labels/brain?_labels.mnc ${SCRATCH}/maget/input/atlases/labels
# copy t2 atlases
cp /project/m/mchakrav/atlases_from_git/T2_extracted/brain?_t2.mnc ${SCRATCH}/maget/input/atlases/brains/
# rename t2 atlases
rename _t2.mnc .mnc ${SCRATCH}/maget/input/atlases/brains/*.mnc
Quality control
The best way to QC is to overlay each subjects' labels onto their input file in Display. Download the labels (${SCRATCH}/maget/output/fusion/majority_vote/*.mnc) and images (${SCRATCH}/maget/input/subjects/brains/*.mnc) to your computer and examine them:
Display -gray ABC_101_T2_corrected.mnc -label ABC_101_T2_corrected_labels.mnc
You can write a loop to go through all subjects by creating a list of files to loop through, for example:
#!/bin/bash
#Loop through MAGeT outputs based on subjects list from text file and open in Display for QC.
source /opt/minc/1.9.17/minc-toolkit-config.sh
while read subj; do
file=${subj}*.cutneckapplyautocrop.beastextract.mnc
labels=${subj}*.cutneckapplyautocrop.beastextract_labels.mnc
if [ -f ${file} ];then
Display -gray ${file} -label ${labels}
fi
done < files.txt
For very large datasets, where this is not feasible, you can generate QC images. Here is one example of how to do it on the CIC:
#!/bin/bash
dir="/path/to/maget/dir"
imgdir="/path/to/inputs"
labeldir="/path/to/output/fusion/majority_vote"
mkdir -p ${dir}/QC
for subj in `ls ${imgdir}`; do
echo ${subj%.*}
if [ -e ${labeldir}/${subj%.*}_labels.mnc ] && ! [ -e ${dir}/QC/${subj%.*}_QC.jpg ]; then
echo MAGeT-QC.sh ${imgdir}/${subj} ${labeldir}/${subj%.*}_labels.mnc ${dir}/QC/${subj%.*}_QC.jpg >> magetQCjob.txt
elif [ ! -e ${labeldir}/${subj%.*}_labels.mnc ]; then
echo ${subj} >> ${dir}/missing_maget_labels.txt
fi
done
Then run your script on the CIC as follows:
module load qbatch minc-toolkit/1.9.18 minc-toolkit-extras/1.0
qbatch -N QC --chunksize 1 --walltime=0:10:00 magetQCjob.txt
Note. This will take some time if you have many subjects. For LAM high resolution T2-weighted images, there is a bug, where it fails on some subjects and shows a mincresample usage/help output. I've found that running it multiple times (the above script will skip already completed files) ends up working eventually, but upcoming new version of minc-toolkit may help.
Rows 1 & 2 are coronal slices, 3 & 4 are saggital, 5 & 6 are axial. Odd numbered rows are without the labels and even numbered rows are identical to the row above it but with the labels visible. From left to right columns, coronal images move from posterior to anterior, saggital images move from left to right, and axial images move from inferior to superior. Go carefully through these images (zoom in) and rate on a 0,1,2 scale:
- 0 (FAIL): label is missing or in the wrong place, large or many small errors
- 1 (SMALL ERRORS): slight over or under segmentation in a few slices
- 2 (PERFECT/NEAR PERFECT): labels cover structure exclusively, look good in (almost) all slices
Colour coding for the hippocampal and white matter labels in Display are as follows:
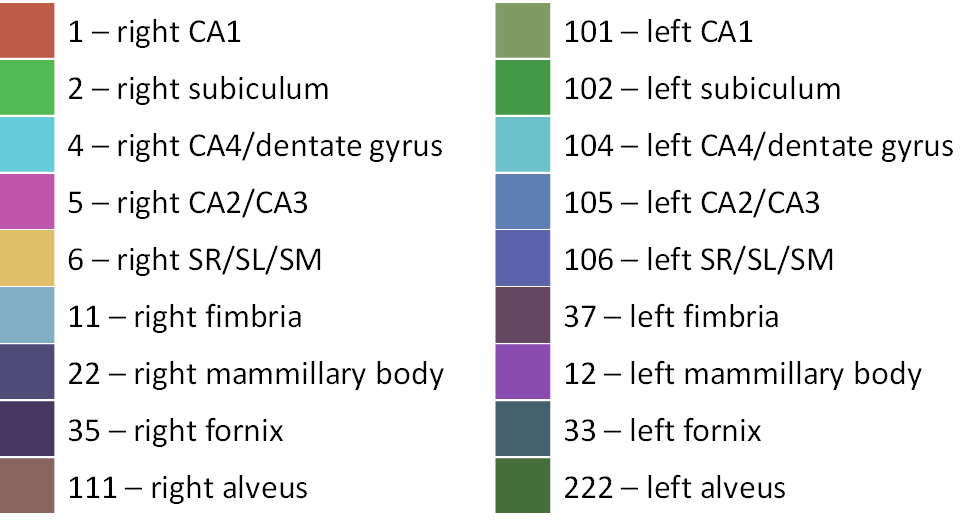
Running MAGeTbrain updates
For ongoing studies, you may want to run MAGeT on only updated subjects. This can be done if you saved certain files from the original analysis, notably the templates and the atlas-to-template registrations, which are located in output/registrations. Copy these back to Niagara and put them into the template and output/registrations directory once you have run mb init. It will take quite some time to copy the registrations back and forth, so if you have a project allocation on Niagara, it would be better to store them there. You will also want to make sure you backup the version of MAGeT you were using and use that one instead of downloading it again from GitHub.