Home - CDCgov/HantaNet GitHub Wiki

Welcome to the HantaNet wiki! HantaNet is a standalone Hantavirus data visualization engine, powered by MicrobeTrace. This site shows you a specific use-case scenario for MicrobeTrace, which is a free, secure web-browser based visualization tool that facilitates the integration and analysis of laboratory, epidemiological and sequence data.
Hantaviruses are RNA viruses with a segmented genome that are shed via urine, feces, and saliva from an infected animal host and spread to people via aerosolized virus-containing particles. Each hantavirus strain is predominantly associated with one rodent species or few closely related species. Hantavirus Pulmonary Syndrome (HPS) is a respiratory disease caused by "New World" hantaviruses (e.g. Sin Nombre Virus) and Hemorrhagic Fever with Renal Syndrome (HFRS) is caused by "Old World" hantaviruses (e.g. Seoul Virus). Studies have shown that "New World" and "Old World" hantaviruses are phylogenetically distinct.
Please check out our publication titled HantaNet: A New MicrobeTrace Application for Hantavirus Classification, Genomic Surveillance, Epidemiology and Outbreak Investigations. We also have a HantaNet Tool Demonstration Video available.
Polar Expedition Cruise Ship MV Hondius Andes Hantavirus Outbreak (April–May 2026)
As of May 11, 2026, there are four genome sequences available in Pathoplexus.org (accessions PP_006WBLH.1, P_006W3U9.1, PP_006W6RC.2, PP_006WANE.1) under restricted use. The Hodcroft Lab has built phylogenies of the three gene segments in Nextstrain.org including the outbreak sequences. A link to MicrobeTrace 2.0 is available under the "View in other platforms" tab at the bottom of the built page. The network below was generated using MicrobeTrace from the hantavirus S segment tree and was modified to show the minimum spanning tree, with recent Andes virus sequences from the outbreak highlighted as green nodes and overlaid on a map.
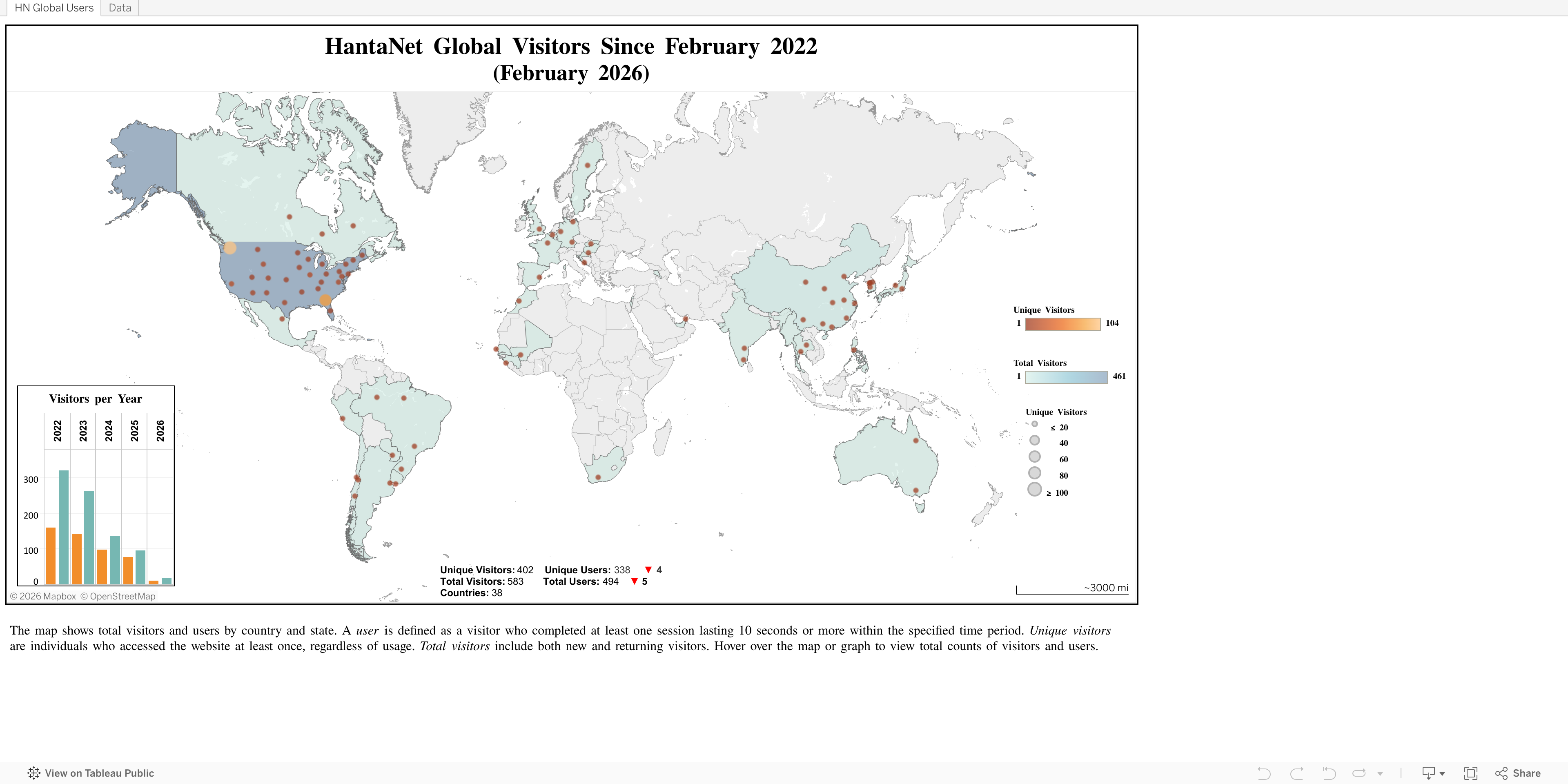
HantaNet Global Visitors
As of May 11, 2026, HantaNet has received more than 600 visitors, including over 500 active users and 800 site visits. During the first two weeks of May, the platform experienced a surge in activity from users in more than 70 countries, likely driven by the ongoing Andes hantavirus outbreak, which began in April 2026. The bar chart shows annual visitor counts from 2022 through 2026. Click the image to open the published dashboard in Tableau Public and explore total counts of visitors and users.
Scope: Web platform for hantavirus data analysis and visualization to improve domestic hantavirus surveillance, standardize and modernize diagnostics, case reporting and data sharing between the CDC and external partners.
HantaNet features:
- Pre-loaded sequence reference sets for each segment (Small, Medium and Large)
- Open-reading frame; stop codon removed
- Collection dates of reference sequences: 1982-2017 (currently loading new Andes outbreak sequences)
- Datasets contain viral sequences of animal and human origin
What you can do:
- Subtype and classify your hantavirus sequences
- Network or phylogenetic tree visualization of your data
- Overlay molecular and epidemiological links
- Visualize spatiotemporal distribution of hantaviruses
HantaNet Project Contacts: Roxana Cintron ([email protected]), Shannon Whitmer ([email protected]), Joel Montgomery ([email protected]), Bill Switzer ([email protected])